Sequence
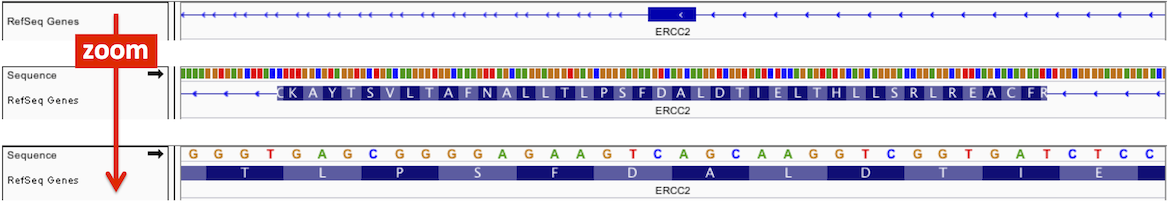
When zoomed in sufficiently, the reference genome Sequence track appears. The sequence is represented by colored bars or colored letters, depending on zoom level, with adenine (A) in green, cytosine (C) in blue, guanine (G) in yellow, and thymine (T) in red.
- You can change the nucleotide colors by selecting View Preferences > General and clicking on the color swatches in the Nucleotide Colors section.
IGV displays the sequence of bases as they appear in the FASTA file for the reference genome. In addition to the upper case letters A, C, G, and T, you may see lower case letters for these bases, and also N/n. Lower case letters often mark repeated regions, and N/n may represent ambiguous nucleotides. However, the convention for the use of case and N, is not completely standardized, and depends on the creator of the genome sequence.
Flipping the strand#
You can change the strand that is displayed by clicking on the arrow in the title to the left of the track. Note that the sequence and the arrow are only displayed when zoomed in to a sufficiently small region.
- Alternatively, right-click on Sequence track to select Flip strand from the pop-up menu.

The direction of the arrow indicates which strand is currently displayed. An arrow pointing left indicates that the negative strand is showing. This strand will show the complement nucleotides and reverse complement translations.
Sequence translation#
With the reference genome sequence track, you can optionally display a 3-band track that shows a 3-frame translation of the amino acid sequence for the corresponding nucleotide sequence. The translation is shown for the strand indicated.
- Right-click on Sequence track to select Show translation from the pop-up menu and to select a Translation Table.

Amino acids are displayed as blocks colored in alternating shades of gray. Methionines are colored green, and all stop codons are colored red. When you zoom all the way in, the amino acid symbols will appear.
You can toggle the display of this translation track by clicking once, anywhere in the sequence or translation track, or by toggling Show Translation in the track popup menu.