Sample Information
Sample information files are tab-delimited text files used to associate attributes with sample names. The attributes are displayed in the "Sample Information" panel and can be used to sort, group, and filter tracks.
The file consists of up to three sections
- A tab delimited table specifying attributes and their values for each sample - Required.
- Sample mapping information - Optional, needed only if the sample names in the attribute information do not match the sample names in the data files.
- Colors - Optional, used to explicitly specify colors.
Attributes#
The attributes section is denoted by the line #sampleTable. It consists of a tab delimited header row defining the
attribute names, followed by one row per sample containing attribute values. The first column contains the sample names.
The #sampleTable line can be omitted if the attribute section is the first or only section in the file.
An example attribute section for 2 samples follows.
#sampleTable
ID Subtype sil_width GENDER KarnScore Censured MGMT_methylated % Tumor Nuclei % Necrosis
TCGA-02-0001 Classical -0.135526414 FEMALE 80 0 97.5 0
TCGA-02-0002 Neural -0.069669747 MALE NA NA No NA DEAD 0 97.5 5
Sample mapping#
Optional section. The sample mapping section begins with the line #sampleMapping. It is used to map sample names in the sample
information file to corresponding names in the data files. If the names match this section can be omitted.
The format is two-column tab delimited. The first column is
the sample name in the data file; the second column is the sample identifier in the attributes information.
Example:
#sampleMapping
TRIBE_p_TCGAaffx_B1_2_GBM_Nsp_GenomeWideSNP_6_A01_155716 TCGA-02-0001
TRIBE_p_TCGAaffx_B1_2_GBM_Nsp_GenomeWideSNP_6_A03_155748 TCGA-02-0002
Attribute colors#
Optional section. By default, IGV randomly assigns colors to the attribute values. You can optionally specify the colors for attribute values in RGB format for a specific attribute name, a specific value, or as a heatmap scale for numeric columns in monocolor or in two-color heatmap for specified ranges.
The attribute colors file (or section in a combined file) begins with the line #colors. The file is tab delimited
with three or four columns:
- 1: Attribute name. An asterisk
*indicates the color specification applies to all attributes. - 2: Attribute value or range of two values separated by a colon
:. An asterisk*indicates the color specification applies to all attribute values. - 3: Color in RGB format. If a color is also specified in column 4, this is the first color of a two color heatmap.
- 4: (Optional) Second color (RGB format) of a two-color heatmap.
#colors
# A value of "MALE" for the "GENDER" column gets the color (0,0,155)
GENDER MALE 0,0,155
# A value of "Classical" in any column gets the color (80,180,80)
* Classical 80,180,80
# Numeric column example, monocolor heatmap
KarnScore * 0,0,255
# Another monocolor heatmap, this time with the range specified
% Tumor Nuclei 90:100 0,0,255
# A two-color heatmap with the range specified
sil_width -0.1:0.5 0,0,255 255,0,0
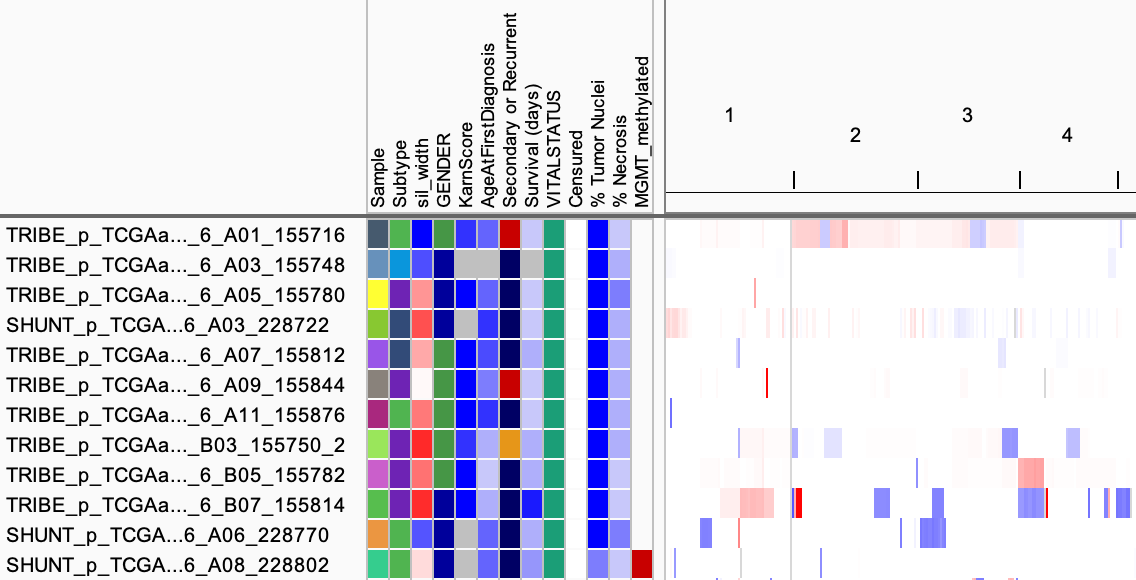
The above example snippets are all from this sample info file, which has
also been used in IGV in the following screenshot.